The headline and subheader tells us what you're offering, and the form header closes the deal. Over here you can explain why your offer is so great it's worth filling out a form for.
Remember:
Interest in artificial intelligence applications for image analysis in healthcare is increasing fast. The number of publications in academic journals concerning machine learning is growing exponentially and consequently several image-recognition applications have been implemented in daily clinical practice. Accurate algorithms can serve as additional diagnostic tools for physicians in order to facilitate and accelerate the workflow and, most importantly, improve the accuracy of the diagnostic process.
 Image analysis algorithms should be added to the physician’s toolbox and not be regarded as a potential replacement of medical specialists. Despite physicians being concerned with and responsible for their patients’ well-being, most are not familiar with the mathematical details of machine learning algorithms. Therefore, medical applications should not serve as ‘black boxes’, but provide additional information on how the model came to its predictions. This blog demonstrates two possible techniques for acquiring this information: probabilistic layers, which can be used to address the uncertainty on a model’s predictions and Grad-CAM a method for demonstrating which pixels contribute most to the model’s outcome. In this blog we will focus on dermatoscopic images, but the presented techniques are applicable to other medical fields (e.g. radiology and pathology) as well.
Image analysis algorithms should be added to the physician’s toolbox and not be regarded as a potential replacement of medical specialists. Despite physicians being concerned with and responsible for their patients’ well-being, most are not familiar with the mathematical details of machine learning algorithms. Therefore, medical applications should not serve as ‘black boxes’, but provide additional information on how the model came to its predictions. This blog demonstrates two possible techniques for acquiring this information: probabilistic layers, which can be used to address the uncertainty on a model’s predictions and Grad-CAM a method for demonstrating which pixels contribute most to the model’s outcome. In this blog we will focus on dermatoscopic images, but the presented techniques are applicable to other medical fields (e.g. radiology and pathology) as well.
The two techniques will be illustrated using Kaggle’s public HAM (“Human Against Machine”) 10000 dataset which contains over 10000 images of 7 classes of pigmented skin lesions, including several types of skin cancer (i.e. melanoma and basal cell carcinoma) and less invasive types like benign keratosis-like lesions and physiological melanocytic lesions.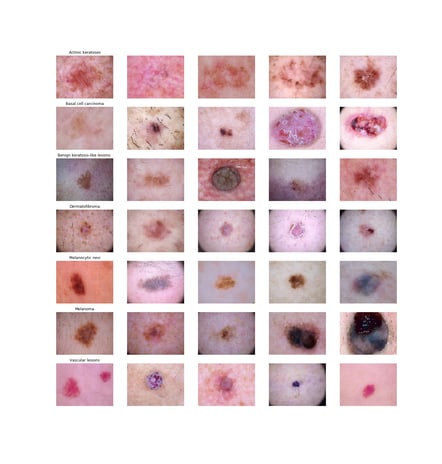 Although the dataset contains additional features as age and location valuable for classifying skin lesions, for illustrative purposes this blog will merely focus on the imaging data. On the left several example images, all obtained with a dermatoscope are depicted.
Although the dataset contains additional features as age and location valuable for classifying skin lesions, for illustrative purposes this blog will merely focus on the imaging data. On the left several example images, all obtained with a dermatoscope are depicted.
First, a basic convolutional neural network was trained using the VGG16 architecture with predefined weights. The weights of the first layers were frozen and only the last 6 layers were retrained. An additional dense layer with a softmax activation function was added.
After 10 epochs of training the transfer learned model yielded a validation accuracy of 82%. At this moment, the output of the model consists of an array representing the probabilities an image belongs to each of the 7 classes (see below).
[[2.3303068e-09 1.2782693e-10 1.4191615e-02 1.8526940e-14 3.9408263e-02 9.4640017e-01 2.1815825e-11]]These predictions can already guide physicians in the right direction, however additional information of the model’s method of ‘thinking’ will increase understanding of and trust in these kind of deep learning models. The next sections will illustrate that for this purpose probabilistic layers and Grad-CAM can be extremely valuable.
Adding a probabilistic layer to a model’s architecture is a valuable method to evaluate the uncertainty of a model’s predictions. The fundamental difference between ‘normal’ and probabilistic layers is that probabilistic layers assume weights to have a certain distribution instead of being point estimates. Probabilistic layers approximate the posterior distribution of the weights by variational inference. Essentially, the posterior distribution is approximated through a Gaussian distributed surrogate function. During each forward pass, weights of a probabilistic layer are sampled from this surrogate distribution.
As a consequence, introducing the same input multiple times will result in slightly different predictions each time. In a classification problem, the model will yield a distribution of probabilities for each class. This probability distribution can be used to asses the uncertainty of a model’s predictions: if the uncertainty is limited, repeated predictions will be very close to each other, while a wide distribution of predictions imply a large model uncertainty.
This blog will focus on implementation of a specific probabilistic dense layer (Flipout), which can be easily achieved using the Tensorflow Probability module. Instead of adding a normal dense layer to the predefined VGG16 network we add a dense Flipout layer using the same method. The full mathematical background of Flipout was described by Wen et al. and can be found at https://arxiv.org/abs/1803.04386.
After training the model, the output of our dense Flipout layer looks similar at first glance: an array representing a probability for each of the 6 classes. However, as aforementioned each prediction of the same image is slightly different as a result of the Flipout layer sampling weights out of a distribution each forward pass. By repeating the predictions for a certain image multiple times, a distribution of probabilities can be obtained for each of the 7 classes of skin lesions.
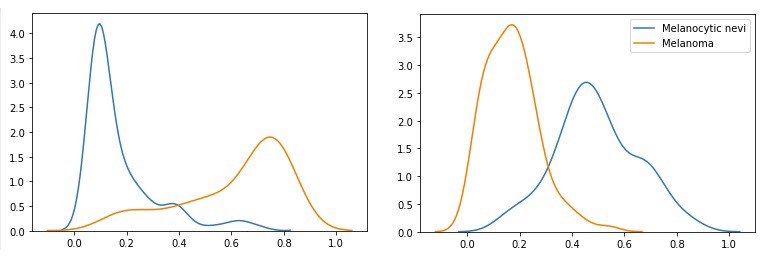
Density plots of two images passing through the network 100 times
Each of the images on the left shows the probability distribution of an image belonging to the class ‘melanocytic nevi’ (blue) or ‘melanoma (orange) generated by passing each image through the network 100 times. The x-axis reflects the probability of the image belonging to one of the classes, while the height of the plots express the distribution density.
The value of the additional information for physicians can be illustrated with the depicted density plots. Let’s have a look at the left plot: passing the first image through a ‘normal’ convolution neural network would yield a low point estimate for the class ‘melanocytic nevi’ (blue) and a high point estimate for the class ‘melanoma’ (orange). The probabilistic Flipout layer enables the evaluation of the uncertainty of these predictions. Specifically, the left plot shows little variation in the predicted (low) probabilities for the class ‘melanocytic nevi’. However, the distribution of the ‘melanoma’ class has much more variation, implying that the probability of the image belonging to the class ‘melanoma’ is quite uncertain.
In summary, a ‘normal’ convolutional neural network would provide a probability for each class, while a model containing a probabilistic Flipout layer yields a distribution of probabilities for each class and can thus be used to evaluate the uncertainty of the model’s predictions. Physicians can use this extra information to gain a better understanding of the predictions and use his/her knowledge to confirm or exclude a certain diagnosis.
Gradient-weighted Class Activation Mapping (Grad-CAM) is a method that extracts gradients from a convolutional neural network’s final convolutional layer and uses this information to highlight regions most responsible for the predicted probability the image belongs to a predefined class.
The steps of Grad-CAM include extracting the gradients with subsequent global average pooling. A ReLU activation function is added to only depict the regions of the image that have a positive contribution to the predefined class. The resulting attention map can be plotted over the original image and can be interpreted as a visual tool for identifying regions the model ‘looks at’ to predict if an image belongs to a specific class. Readers interested in the mathematical theory behind Grad-CAM are encouraged to read the paper by Selvaraju et al. via https://arxiv.org/abs/1610.02391.
The code below demonstrates the relative ease of implementing Grad-CAM with our basic model.
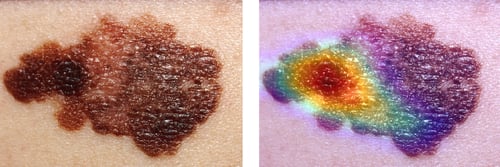
These images demonstrate the original image of a melanoma and the same image with a superimposed attention map created using the Grad-CAM algorithm. These images clearly illustrate that Grad-CAM yields additional information that is useful for daily clinical practice. This attention map reflects which parts of the skin lesion is affecting the model’s predictions most. Possibly, this information can be valuable to guide skin biopsies in search for confirmation of the suspected diagnosis. In addition, provided that the algorithm is accurate in it’s predictions, repeated use of the algorithm with incorporated visual features in different patients will increase a physician’s trust in the algorithm’s predictions.
Convolutional neural networks have the potential to provide additional diagnostic tools for several medical specialties as radiologists, dermatologists and pathologists. However, models that merely provide class-predictions are not sufficient. We believe that introducing methods as adding a probabilistic reasoning and Grad-CAM provide insight in the model’s decisive properties and are therefore essential for physicians to understand and start embracing machine learning algorithms in daily clinical practice.
Special thanks to Richard Bartels and Loek Gerrits
Lorem ipsum dolor sit amet, consectetur adipiscing elit

These Stories on Data Science
Coltbaan 4C
3439 NG NIEUWEGEIN
+31 30 227 2961
info@vantage-ai.com
Follow us: